 Intracellular compartmentalisation of metabolism & functions
Intracellular compartmentalisation of metabolism & functions
Partner Institutes : IBS, BIG
Objectives:
A characteristic of many cellular processes is their spatial compartmentalization. Most cellular events do not occur ubiquitously throughout the cell but are limited to specific, spatially defined sites that often occur in dedicated cell structures. Remarkably, common mechanisms appear to organize some of the vastly different, fundamental cellular processes. The aims of this project are to decipher the molecular mechanisms underlying the regulation of the chloroplast biogenesis and function and to expand our understanding of the processes underlying the adaptation of the chloroplast to various stress conditions.
Background:
The chloroplast is an organelle that assimilates atmospheric inorganic carbon, reduce and assimilate nitrogen and sulfur, but is also a unique cell compartment, being involved in the partial or complete synthesis of a huge number of metabolites (lipids, pigments, sugars, amino acids, vitamins….) that are essential for animal feeding. For this reason, the chloroplast is considered as an essential element of the food chain supporting a large part of life on the earth, energy …
The chloroplast is a model of choice to elaborate a quantitative and dynamic model of a metabolic system. On a practical point of view, the total number of enzymatic reactions is expected to reach 1000 in the chloroplast, when considering the total number of chloroplast proteins (around 3,000). The functional complexity of the chloroplast is thus lower than that of model bacteria like Escherichia coli. Some authors have already demonstrated that the bacterial metabolism can be defined in details, and in a comprehensive way, at the level of a complete system (http://gcrg.ucsd.edu/organisms/ecoli.html). This “relative simplicity” suggests that a community of scientists working on chloroplast structure and function should be able, at the scale of a decade, to produce more and more precise and predictive models of the metabolic comportment of an entire chloroplast, and then of the dynamics of this system.
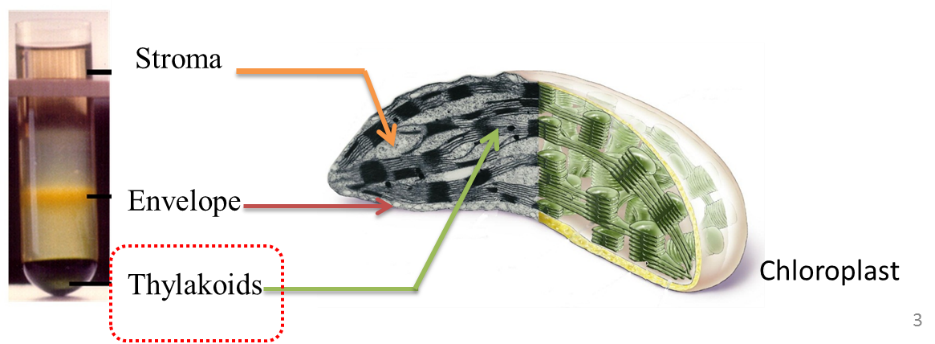
The main limit of such an approach results from the heterogeneity of the produced data and of the diversity of the addressed questions. We thus propose to start the project with defined specific plastid metabolisms or functions that make the core of the analyses. A true integration is thus required and the characterization of the system will be performed in a reference state allowing the entire community to analyze the data in a quantitative point of view. We also propose to use the entire imaging technologies available in the three institutes to produce a physical and virtual model of the chloroplast at all scales of biology. Finally, the dynamics of this compartment will also be analyzed, thus allowing to propose predictive mechanistic models of the chloroplast functioning and adaptation to various environmental constraints.
Expected results and perspectives
- Towards the virtual chloroplast : Integration of all data (including structural data) in a specific database containing information on chloroplast structure and functions at all scales

- New concepts of spatiotemporal organization to optimize metabolic channelling, introduce new concepts for membrane lipid metabolism with shape constraints, settle in functions to limited space within the organelle, protect cell components from ROS produced during photosynthesis.
- Setup of new analytical strategies and new methodologies that will be adapted to the study of other biological models. The generation of knowledge and the integration of complementary approaches to address a biological question are attractive for the scientific community and will lead to important dynamics within the three local institutes.
- Towards synthetic biology: optimization of channelling. Exploitation of data for downstream applications aiming to forecast changes in chloroplast metabolic functions for applications like the production of biofuels and molecules issued from the diversity of the secondary metabolism: they can be valued as surrogates for petrochemistry (precursors for plastics and other industrial polymers).
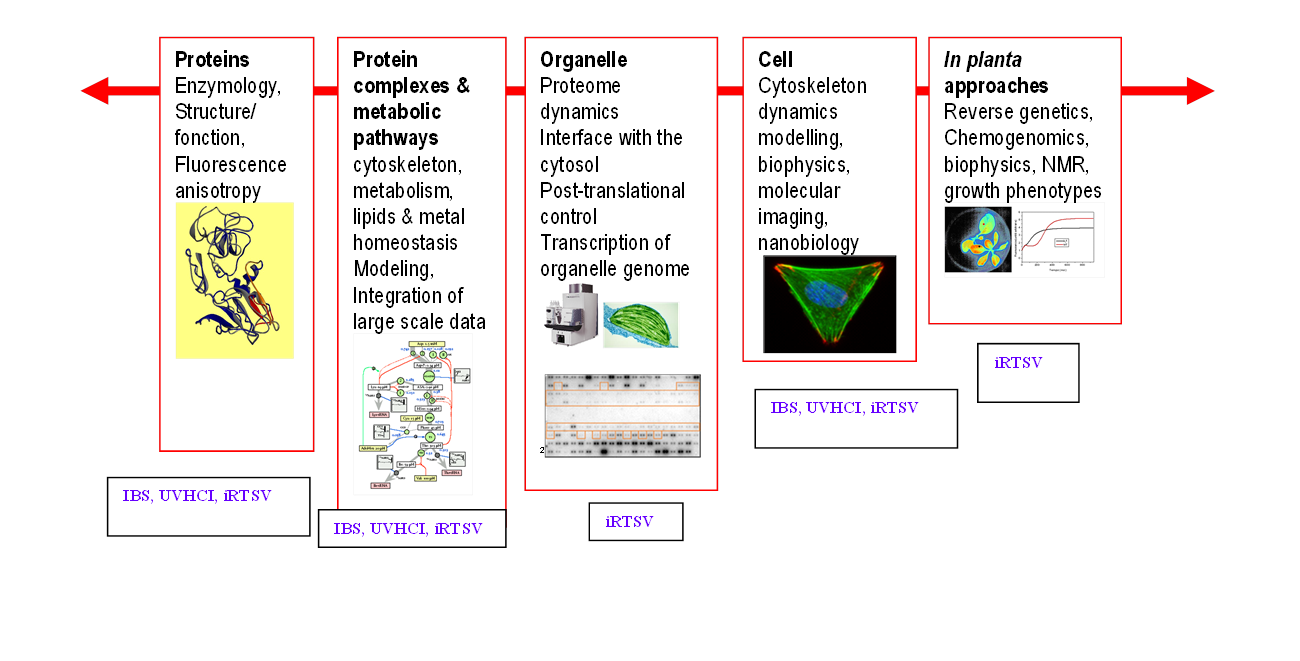
Figure : Research on chloroplast at different levels of integration and the involvement of the different GRAL teams in the project.
