C.Petosa, IBS : Nuclear export of a viral ribonucleoprotein
Objectives:
This project aims to: (1) determine the stoichiometry and assembly pathway of the RRE/Rev/CRM1/Ran/RanBp3 export complex; and (2) determine the 3D architecture of this export complex.
Background:
An important step in the HIV life cycle is the export of incompletely spliced viral transcripts from the nucleus to the cytosol. A key player in this process is the HIV protein Rev. In the nucleus Rev recognizes the viral RNA genome and other unspliced viral transcripts by binding a 350-nucleotide Rev-response element (RRE). These transcripts are then exported to the cytosol by host cell proteins recruited by Rev, including the export factor CRM1, the small GTPase Ran and Ran-binding protein 3 (RanBP3). Interfering with Rev/RRE export is a potentially attractive anti-retroviral therapeutic strategy.
Methodology:
The RRE/Rev/CRM1/Ran/RanBp3 holo-complex and sub-complexes will be assembled by incubating WT and mutant constructs of Rev, CRM1, Ran and RanBP3 with in vitro transcribed RRE and partial RRE fragments. The stoichiometry of these complexes will then be determined by biophysical techniques (native mass spectrometry, analytical ultracentrifugation, dynamic and multi-angle light scattering) while molecular shape information will be determined by small-angle scattering (SAXS/SANS) and negative-stain electron microscopy. Results will be combined with known crystal structures of CRM1, Ran, Rev and RanBP3 to deduce a three-dimensional architecture of the holo-complex.
Team:
Carlo Petosa: Team Leader (CNRS DR2)
Mizar Oliva: PhD Student
Francesca Cosica: PhD Student
Didier Spittler: PhD Student
Fabienne Hans: Maître de Conférences (Université Joseph Fourier)
Marjolaine Noirclerc-Savoye: Research Engineer (CEA)
GRAL recruitment :
GRAL support allowed PhD student Mizar Oliva to be recruited in October 2013 to work on this project.
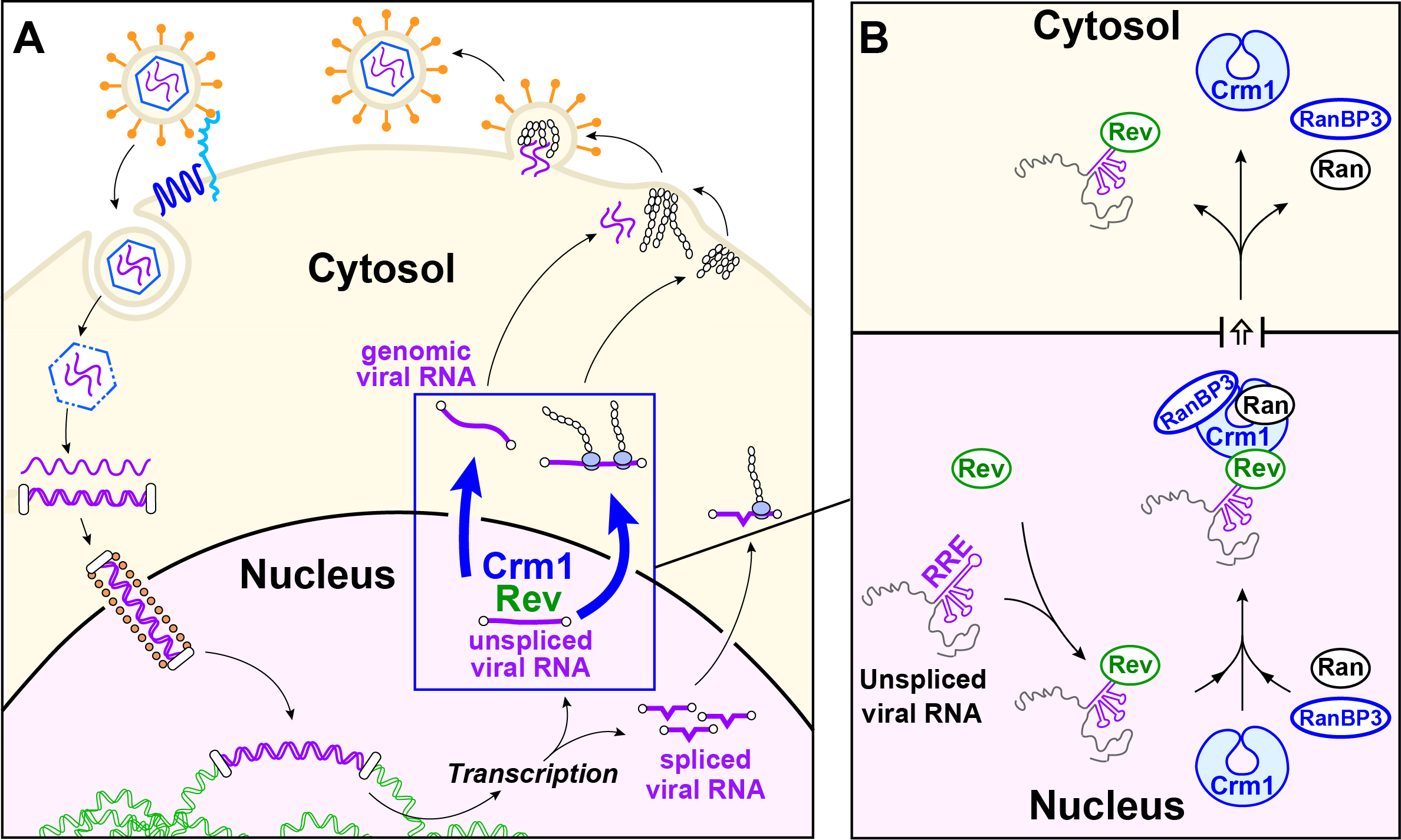
Figure 1. Nuclear export of an HIV-1 ribonucleoprotein. A.Cartoon summarizing the HIV-1 infection cycle. Steps include viral binding to the host cell surface, internalization, release of the viral RNA into the cytosol, reverse transcription, integration of the proviral DNA into the host chromosome, transcription, nuclear export of viral RNA, viral assembly, budding and release of newly generated viruses. B. Export of unspliced HIV-1 RNA. Unspliced viral RNA contains a Rev-Response Element (RRE) that is recognized by HIV-1 protein Rev. The Rev/RRE complex then recruits the host cell factors CRM1, Ran and RanBP3, which allow Rev/RRE to be exported to the cytosol.
Publications:
Langer K, Dian C, Rybin V, Müller CW, Petosa C (2011). Insights into the function of the CRM1 co-factor RanBP3 from the structure of its Ran-binding domain. PLoS ONE. 6(2):e17011.
Dian C, Bernaudat F, Langer K, Oliva MF, Fornerod M, Schoehn G, Müller CW, Petosa C. (2013) Structure of a truncation mutant of the nuclear export factor CRM1 provides insights into the auto-inhibitory role of its C-terminal helix. Structure. 21:1338-1349