Weissenhorn, BIG: Structure of ESCRT-III and HIV-1 budding
Objectives
Provide structural information on ESCRT-III polymers in vitro and in vivo to understand their role in membrane fission.
Background
A large number of enveloped RNA viruses including HIV-1 employ the ESCRT (endosomal sorting complexes for transport) machinery for virus release. This final step in budding is catalyzed by ESCRT-III most likely with help from the AAA-type ATPase VPS4. ESCRT-III proteins are small helical assemblies and their autoinhibited conformation is controlled by the C-terminal region. Release of the autoinhibition activates ESCRT-III proteins, which leads to polymerization on membranes. ESCRT-III polymer formation is thought to be required for ESCRT-catalyzed membrane remodeling processes such as membrane fission. Our works aims to define the structure of ESCRT-III to determine its functional role in membrane fission.
Methodology
We employ a combination of X-ray crystallography, cryo electron microscopy and cellular cryo electron tomography to obtain different levels of structural information on ESCRT-III filament formation.
Results
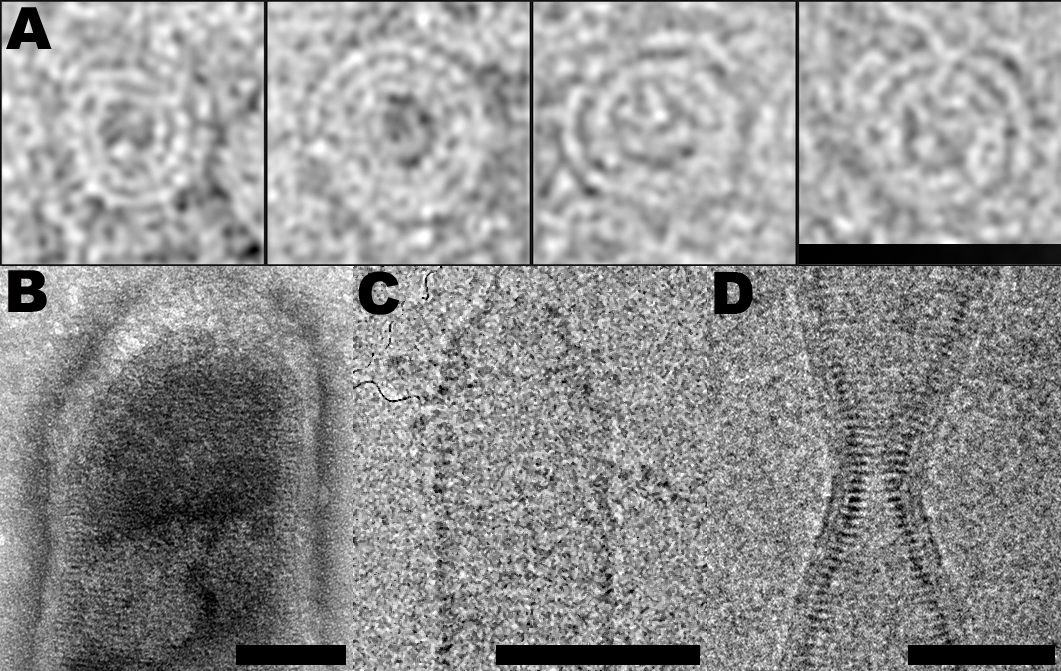
We have previously shown that ESCRT-III CHMP2A-CHMP3 form helical tubular structures. We have now determined the structure of these tubes at medium resolution (22 Å) by electron microscopy. We show that the tube diameters range from 44 to 54 nm and their helical pitches are 30 Å or multiples (n) thereof forming 1-start to n = 6-start helical filament tubes. The structure revealed an elongated 70 Å long asymmetric unit that is tilted by ~ 45º with respect to the tube axis thus producing a 40 Å tube width and a 40 Å external dimension of the asymmetric unit. We also showed that polymer formation rendered the core of the CHMP proteins protease resistant. Furthermore, we demonstrate that CHMP3 exerts an important synergitis effect on HIV-1 budding in the presence of CHMP2A, but not CHMP2B (Effantin et al., 2013).
Team
Several members of the Weissenhorn group contribute to structural biology projects on ESCRT-III including Gregory Effantin, who was supported be a post doctoral fellowship from the ANRS
Gral recruitment
Jens Radzimanowski was recruited in 01/05/2013 (until 30/04/2015) to work on the project.
Publications
Effantin, G., Dordor, A., Sandrin, V., Martinelli, N., Sundquist, W.I., Schoehn, G., and Weissenhorn, W. (2013). ESCRT-III CHMP2A and CHMP3 form variable helical polymers in vitro and act synergistically during HIV-1 budding. Cell Microbiol 15, 213-226
Figure 1: ESCRT-III filaments follow similar structural principles. CHMP2A spirals (negative stain); CHMP2B dome surrounded by membrane (negative stain); CHMP2A-CHMP3 dome (cryo-EM); CHMP2B membranous bottleneck (cryo-EM). Scale bars are 50 nm.
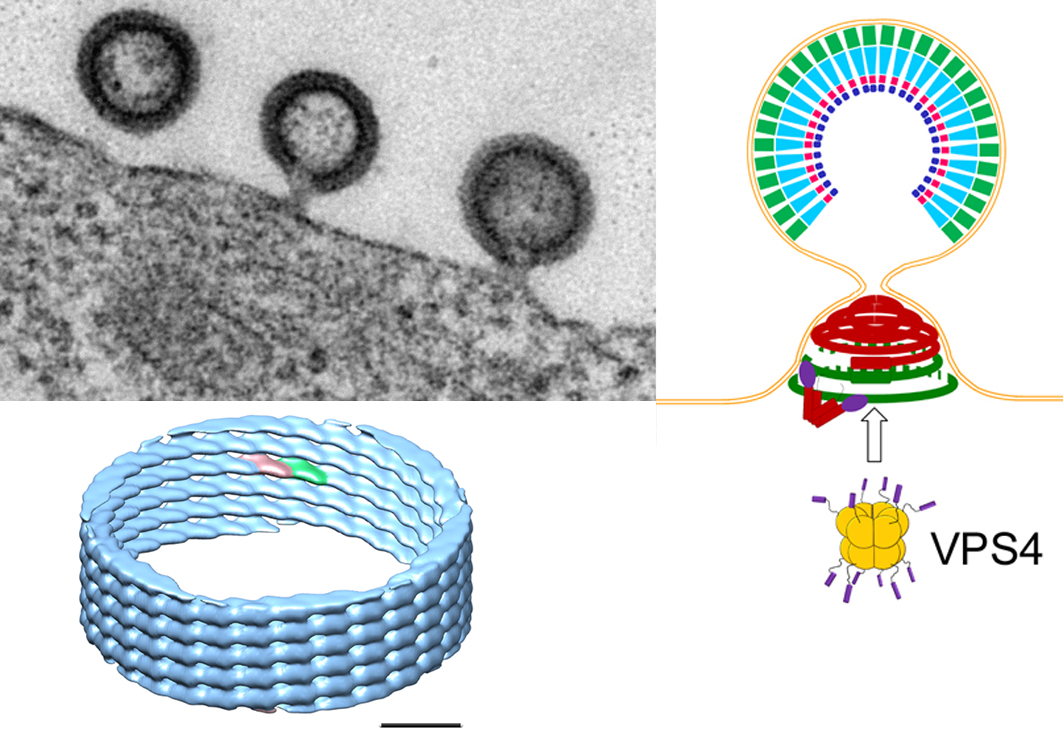
Figure 2: HIV-1 buds from the plasma membrane (upper left panel: EM picture of HIV-1 buds) and recruits ESCRT proteins including ESCRT-III and VPS4. ESCRT-III CHMP2A-CHMP3 form helical polymers in vitro (lower left panel: cryo EM structure at 22 Å resolution) that might assemble inside the neck of the bud and coil up to form dome-like structures (see figure 1). Such structures could attract membrane and thus lead to neck constriction that will favor membrane fission. Additional energy for the final process may be provided by the ATPase VPS4 (upper right panel: model of ESCRT-III induced membrane neck constriction; Gag inside the virion; ESCRT-III polymers in the neck: green CHMP4 filaments coordinated by dimeric Alix (V-shaped) and red CHMP2A-CHMP3 filaments).